VolViewer: Difference between revisions
JeromeAvondo (talk | contribs) No edit summary |
JeromeAvondo (talk | contribs) No edit summary |
||
| (56 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
[[Software| | [[Software#VolViewer| Back to BanghamLab software]]<br> | ||
= | =<span style="color: Navy">What? How? Where?</span>= | ||
[[Image:VolViewer.png|256px|thumb|The VolViewer main application window.]] | [[Image:VolViewer.png|256px|thumb|The VolViewer main application window.]] | ||
VolViewer is | <span style="color: Navy">'''What'''?</span> VolViewer is used for''' viewing volume images''' from, for example, '''confocal''' microscopy or optical projection tomography ('''OPT'''). | ||
Features: | <span style="color: Navy">Features:</span> | ||
* Real-time volume rendering using an optimized 3D texture slicing algorithm. | * Real-time volume rendering using an optimized 3D texture slicing algorithm. | ||
| Line 16: | Line 16: | ||
* Real-time volume clipping. | * Real-time volume clipping. | ||
* 3D measurements, filters & segmentation. | * 3D measurements, filters & segmentation. | ||
* Key frame interpolation for | * Key frame interpolation for movie export. | ||
* Stereo rendering using either quad buffer or anaglyph mode. | * Stereo rendering using either quad buffer or anaglyph mode. | ||
* Scripting interface to other systems, e.g. Matlab, OMERO, etc. | |||
<span style="color: Navy">'''How'''?</span> It is open source and written in C++ using OpenGL, OpenCL and Qt.<br> | |||
<span style="color: Navy">'''Where'''? </span>Binaries are available for the Windows, Mac OS X and Linux, see below. | |||
Requirements: An OpenGL 2.1 / GLSL 1.20 compatible GPU with a recomended 512MB of memory. | Requirements: An OpenGL 2.1 / GLSL 1.20 compatible GPU with a recomended 512MB of memory. | ||
==User Documentation | =<span style="color: Navy">User Documentation</span>= | ||
{| border="1" cellspacing="5" cellpadding="5" style="background-color:#cccccc;" | {| border="1" cellspacing="5" cellpadding="5" style="background-color:#cccccc;" | ||
|- | |- | ||
| align="center"|[[VolViewerUserManual|Quick Guide]] | |||
[[VolViewerUserManual|Quick Guide]] | | align="center"|[[VolViewerCourse|TUTORIALS]] | ||
[[VolViewerTutorials|Video | | align="center"|[[VolViewerTutorials|Video Demos]] | ||
| align="center|[http://dmbi.nbi.bbsrc.ac.uk/index.php/VolViewerScriptsAPI SCRIPTING] | |||
|} | |} | ||
=Sample Data= | =<span style="color: Navy">Sample Data</span>= | ||
{| border="0" cellspacing="5" cellpadding="5" style="background-color:#ffffff; text-align:center;" | {| border="0" cellspacing="5" cellpadding="5" style="background-color:#ffffff; text-align:center;" | ||
| Line 46: | Line 44: | ||
| align="center"| '''Antirinhium Meristem''' || align="center"| '''Arabidopsis Seedling''' || align="center"| '''Arabidopsis Leaf''' <small>(GL2:GUS expression in red)</small> || align="center"| '''Arabidopsis Leaf''' <small>(Ath8:::GUS expression in red)</small> | | align="center"| '''Antirinhium Meristem''' || align="center"| '''Arabidopsis Seedling''' || align="center"| '''Arabidopsis Leaf''' <small>(GL2:GUS expression in red)</small> || align="center"| '''Arabidopsis Leaf''' <small>(Ath8:::GUS expression in red)</small> | ||
|- | |- | ||
| align="center"| [http:// | | align="center"| [http://cmpdartsvr3.cmp.uea.ac.uk/downloads/papers/PlantCellOPT/Antiriniuhm_Meristem(r512g110usmall).zip Download] || align="center"| [http://cmpdartsvr3.cmp.uea.ac.uk/downloads/papers/PlantCellOPT/Arab_Seedling(174).zip Download]|| align="center"| [http://cmpdartsvr3.cmp.uea.ac.uk/downloads/papers/PlantCellOPT/Arab_LeafGL2_GUS(624).zip Download] || align="center"|[http://cmpdartsvr3.cmp.uea.ac.uk/downloads/papers/PlantCellOPT/ArabidopsisLeafAth8_GUS(460).zip Download] | ||
|} | |} | ||
<nowiki>*</nowiki> all data courtesy of Karen Lee [mailto:kareen.lee@bbsrc.ac.uk] | <nowiki>*</nowiki> all data courtesy of Karen Lee [mailto:kareen.lee@bbsrc.ac.uk] | ||
=Download= | =<span style="color: Navy">Download</span>= | ||
Although we try to keep up to date builds these sometimes lag behind the SVN trunk. So if you want the latest version / features, it is best to build the application from the trunk of the SVN. The build system is based on [http://doc.qt.nokia.com/latest/qmake-manual.html qmake] for easy cross platform compilation. | Although we try to keep up to date builds these sometimes lag behind the SVN trunk. So if you want the latest version / features, it is best to build the application from the trunk of the SVN. The build system is based on [http://doc.qt.nokia.com/latest/qmake-manual.html qmake] for easy cross platform compilation. | ||
==OpenGL + Qt + OpenCL + LibTIFF + OMERO== | |||
{| border="1" cellspacing="5" cellpadding="5" style="background-color:#cccccc;" | {| border="1" cellspacing="5" cellpadding="5" style="background-color:#cccccc;" | ||
|- | |- | ||
| align="center"|[http:// | | align="center"|[http://cmpdartsvr3.cmp.uea.ac.uk/downloads/software/BioptonicsViewerV2/VolViewerInstaller_x86.exe Windows (32bit)] | ||
| align="center"|[http:// | | align="center"|[http://cmpdartsvr3.cmp.uea.ac.uk/downloads/software/BioptonicsViewerV2/VolViewerInstaller_x64.exe Windows (64bit)] | ||
| align="center"|[[VolViewerLinux|Linux]] | | align="center"|[[VolViewerLinux|Linux]] | ||
| align="center"|[http:// | | align="center"|[http://cmpdartsvr3.cmp.uea.ac.uk/downloads/software/VolViewerOpenSource/VolViewer.dmg MacOS X (i386/x86_64/10.5+)] | ||
|} | |} | ||
==OpenGL + Qt + LibTIFF== | |||
*Coming soon. | |||
==Windows Specific Notes== | |||
*You may need to install the corresponding Microsoft Visual C++ 2008 Redistributable Package which can be found here: [http://www.microsoft.com/download/en/details.aspx?id=29 32bit] and [http://www.microsoft.com/download/en/details.aspx?displaylang=en&id=15336 64bit]. | |||
*WindowsXP users will need to change the '''view_gldrawbuffer = "GL_FRONT_AND_BACK"''' to '''view_gldrawbuffer = "GL_BACK"''' in the settings.ini file. | |||
*The binaries are built with OMERO 4.3.4 support. | |||
=Source Code= | =<span style="color: Navy">Source Code</span>= | ||
Public SVN: [https:// | Public SVN: [https://cmpdartsvr3.cmp.uea.ac.uk/banghamlabSVN/VolViewer/ https://cmpdartsvr3.cmp.uea.ac.uk/banghamlabSVN/VolViewer/] | ||
==Building from source== | ==<span style="color: Navy">Building from source</span>== | ||
* [[VolViewerBuildingFromSource|Building VolViewer from source]] | * [[VolViewerBuildingFromSource|Building VolViewer from source]] | ||
=Media/Press= | =<span style="color: Navy">Image Gallery</span>= | ||
{| border="0" style="background-color:#000000;" | |||
|- | |||
|align="center"| | |||
[[Image:1272_wh_rgb.png|128x128px]] | |||
[[Image:Am0front.png|128x128px]] | |||
[[Image:Anthers.PNG|128x128px]] | |||
[[Image:Anti_flower_OPT.png|128x128px]] | |||
[[Image:Antirrhinum_flower_small.png|128x128px]] | |||
[[Image:Ara_seedling_colour.png|128x128px]] | |||
[[Image:Cells.png|128x128px]] | |||
[[Image:Cs0prxz0.png|128x128px]] | |||
[[Image:GL2_GUS.png|128x128px]] | |||
[[Image:Leaf_trichomes.png|128x128px]] | |||
[[Image:Leaf5.png|128x128px]] | |||
[[Image:LFY_GUS_Arabidopsis_inflorescence_512.png|128x128px]] | |||
[[Image:OleosinSeed.png|128x128px]] | |||
[[Image:OPT_Leaf_copy.png|128x128px]] | |||
[[Image:Seedling_copy.png|128x128px]] | |||
[[Image:Senecio_floret_copy.png|128x128px]] | |||
[[Image:Snapdragon_Peloric_mutant.png|128x128px]] | |||
[[Image:Tissue.png|128x128px]] | |||
[[Image:Z9r3j2yx.png|128x128px]] | |||
[[Image:Zeds48ci.png|128x128px]] | |||
[[Image:1896_wh_txr_light.png|128x128px]] | |||
[[Image:Ara_flower.png|128x128px]] | |||
[[Image:Arableaf_ath8_OPT.png|128x128px]] | |||
[[Image:Arableaf_young_ath8_OPT.png|128x128px]] | |||
[[Image:Enhby820.png|128x128px]] | |||
[[Image:Nilleafdev.png|128x128px]] | |||
[[Image:Seedling_OPT.png|128x128px]] | |||
[[Image:Senecioclip.png|128x128px]] | |||
[[Image:Silique.PNG|128x128px]] | |||
|} | |||
=<span style="color: Navy">Media/Press</span>= | |||
VolViewer has appeared in the following: | VolViewer has appeared in the following: | ||
[http://www.amazon.co.uk/Handbook-Plant-Science-Keith-Roberts/dp/0470057238/ref=sr_1_19?s=books&ie=UTF8&qid=1289321357&sr=1-19 Front cover: Handbook of Plant Science] | [http://www.plantcell.org/content/18/9.toc Front cover: The Plant Cell] | [http://www.rms.org.uk/Resources/Royal%20Microscopical%20Society/infocus/Edgar%20article.pdf Royal Microscopical Society: Infocus Magazine] | [http://www.bioptonics.com/Home.htm Bundled with the Bioptonic 3001 scanner: Bioptonics Viewer] | [http://www.guardian.co.uk/science/gallery/2007/sep/04/fruitflybrain#/?picture=330675671&index=1 The Guardian newspaper: 3D Fruit fly] | [http://qt.nokia.com/qt-in-use/ambassadors/project?id=a0F20000006LZ2pEAG Qt Ambassador program] | [http://www.triffidnurseries.co.uk/special3.php Triffid Nurseries website] | [http://www.cell.com/cell_picture_show-plantbio Cell: Online Gallery] | [http://www.amazon.co.uk/Handbook-Plant-Science-Keith-Roberts/dp/0470057238/ref=sr_1_19?s=books&ie=UTF8&qid=1289321357&sr=1-19 Front cover: Handbook of Plant Science] | [http://www.plantcell.org/content/18/9.toc Front cover: The Plant Cell] | [http://www.americanscientist.org/issues/pub/2013/1/3d-carnivorous-plants American Scientist] | [http://www.rms.org.uk/Resources/Royal%20Microscopical%20Society/infocus/Edgar%20article.pdf Royal Microscopical Society: Infocus Magazine] | [http://www.bioptonics.com/Home.htm Bundled with the Bioptonic 3001 scanner: Bioptonics Viewer] | [http://www.dailymail.co.uk/sciencetech/article-2215052/The-complexity-intricacy-Mother-Nature-revealed-incredible-pictures-plants--seen-inside.html The Daily Mail] | [http://www.guardian.co.uk/science/gallery/2007/sep/04/fruitflybrain#/?picture=330675671&index=1 The Guardian newspaper: 3D Fruit fly] | [http://qt.nokia.com/qt-in-use/ambassadors/project?id=a0F20000006LZ2pEAG Qt Ambassador program] | [http://www.triffidnurseries.co.uk/special3.php Triffid Nurseries website] | ||
=Author= | =<span style="color: Navy">Author</span>= | ||
* [[Jerome Avondo| Dr Jerome Avondo]] | * [[Jerome Avondo| Dr Jerome Avondo]] Supported by the BBSRC through UEA Computing School and JIC. | ||
Latest revision as of 15:03, 14 August 2013
What? How? Where?
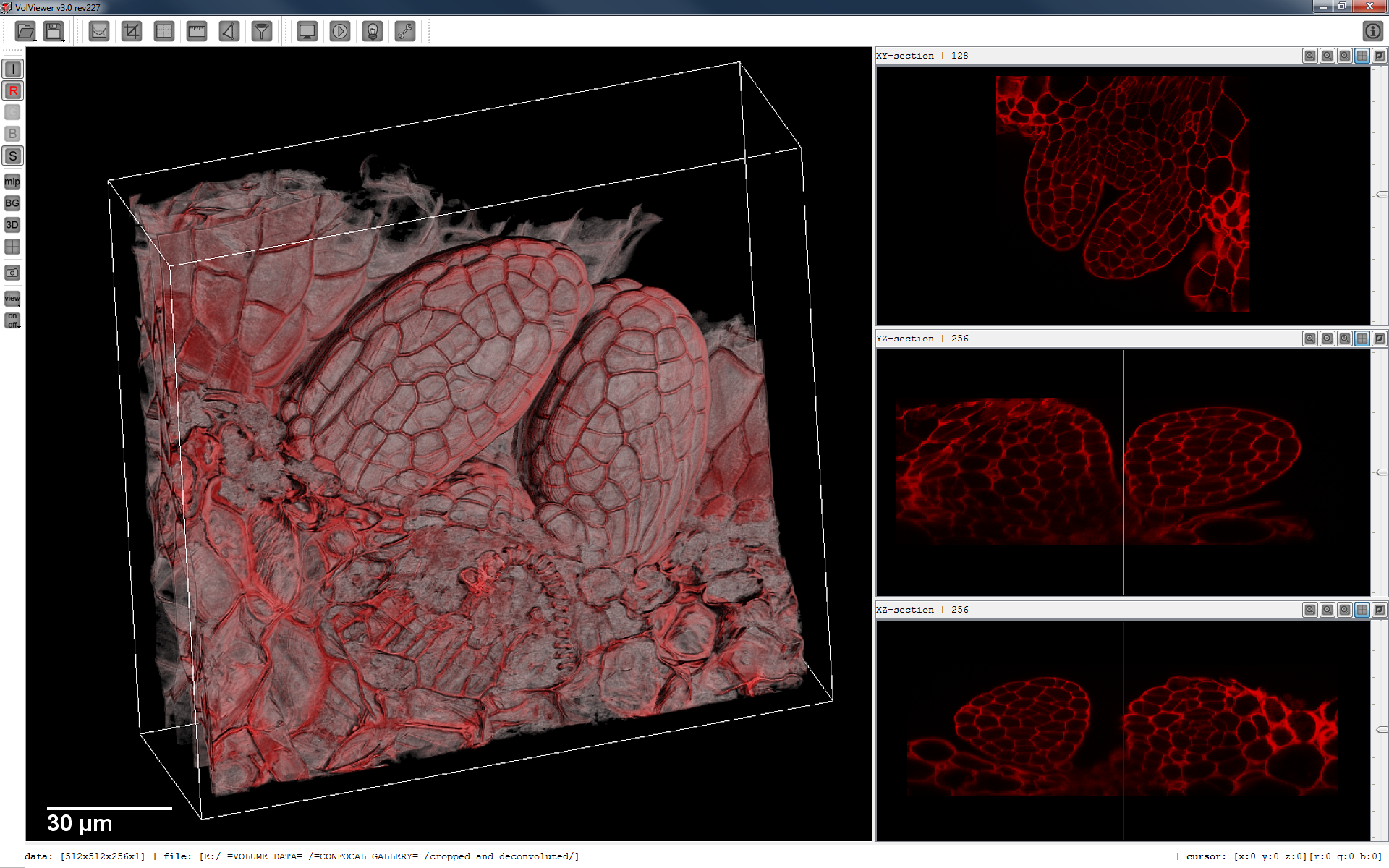
What? VolViewer is used for viewing volume images from, for example, confocal microscopy or optical projection tomography (OPT).
Features:
- Real-time volume rendering using an optimized 3D texture slicing algorithm.
- Interactive transfer functions to independently adjust opacity and intensity for up to three data channels.
- Real-time per channel thresholding, brightness and contrast operators.
- On-the-fly gradient computation for local illumination.
- Iso-surface computation with surface smoothing.
- Section viewing in any orientation / position.
- Real-time volume clipping.
- 3D measurements, filters & segmentation.
- Key frame interpolation for movie export.
- Stereo rendering using either quad buffer or anaglyph mode.
- Scripting interface to other systems, e.g. Matlab, OMERO, etc.
How? It is open source and written in C++ using OpenGL, OpenCL and Qt.
Where? Binaries are available for the Windows, Mac OS X and Linux, see below.
Requirements: An OpenGL 2.1 / GLSL 1.20 compatible GPU with a recomended 512MB of memory.
User Documentation
| Quick Guide | TUTORIALS | Video Demos | SCRIPTING |
Sample Data
 |
 |
 |

|
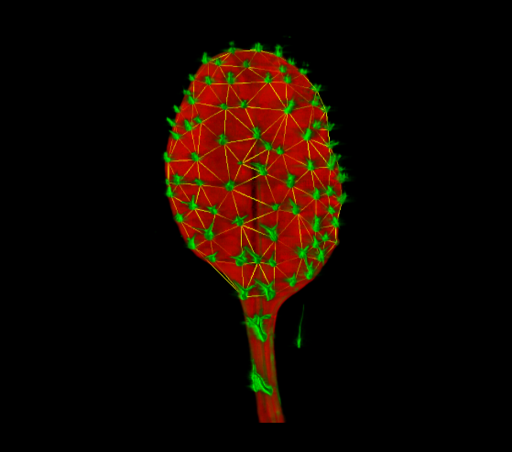
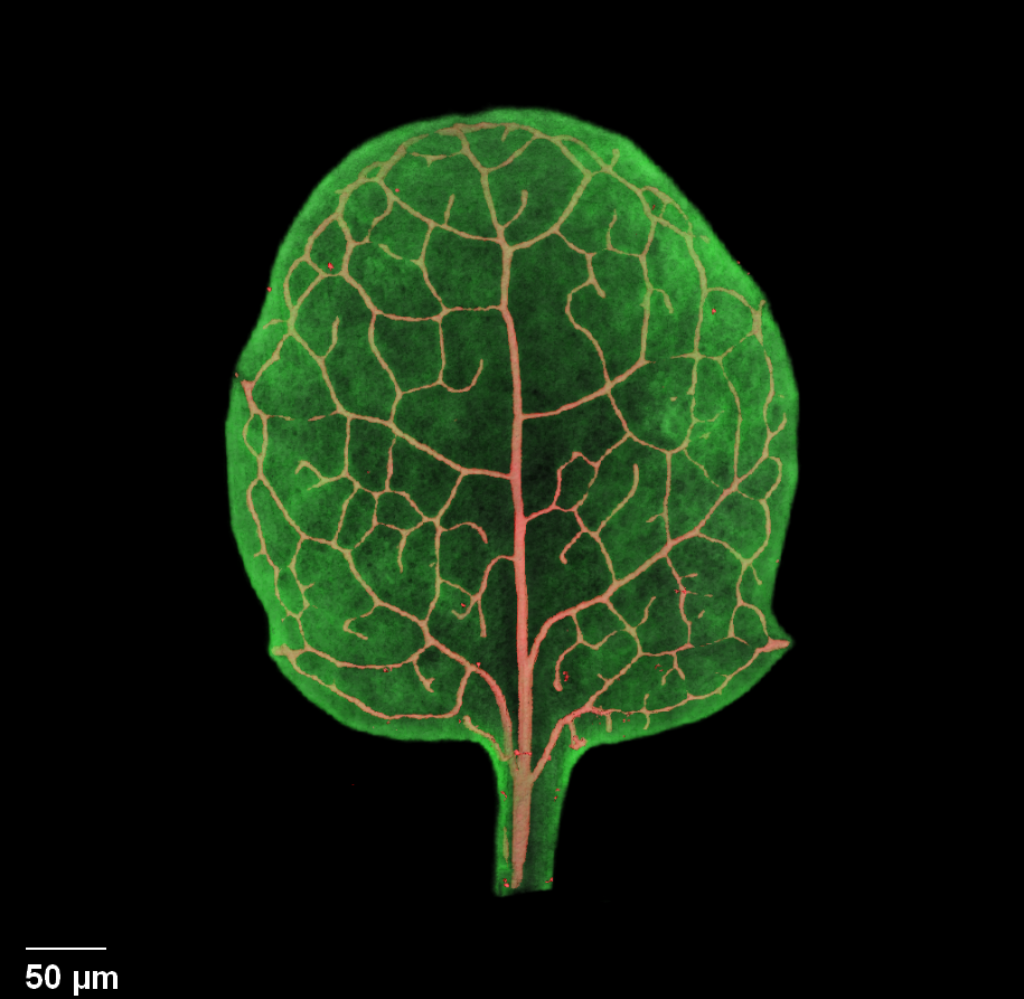
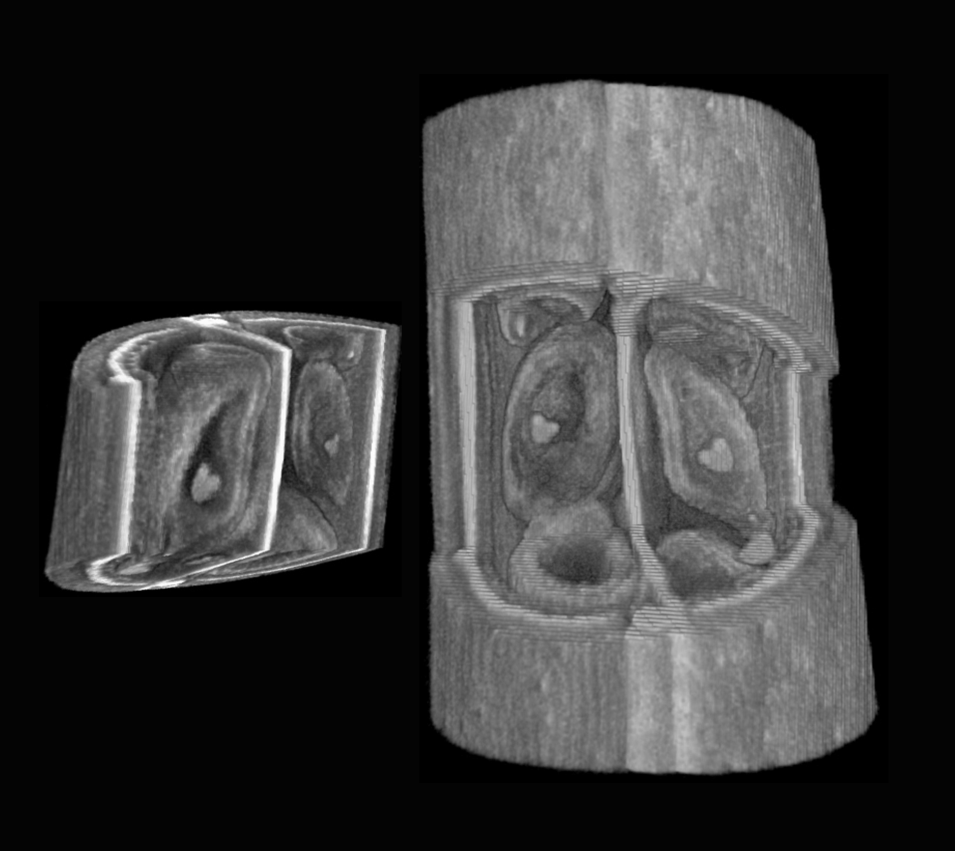
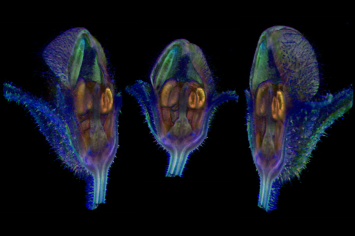
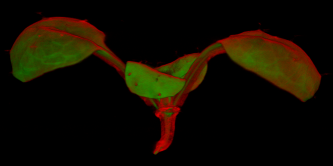
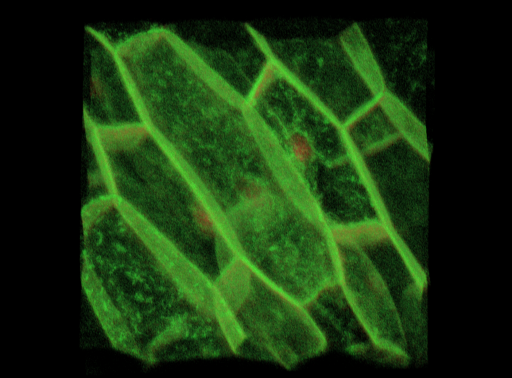
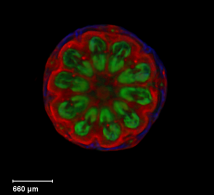
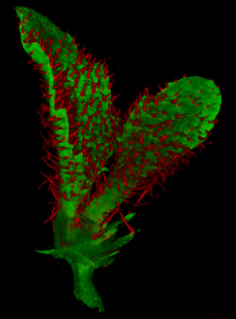
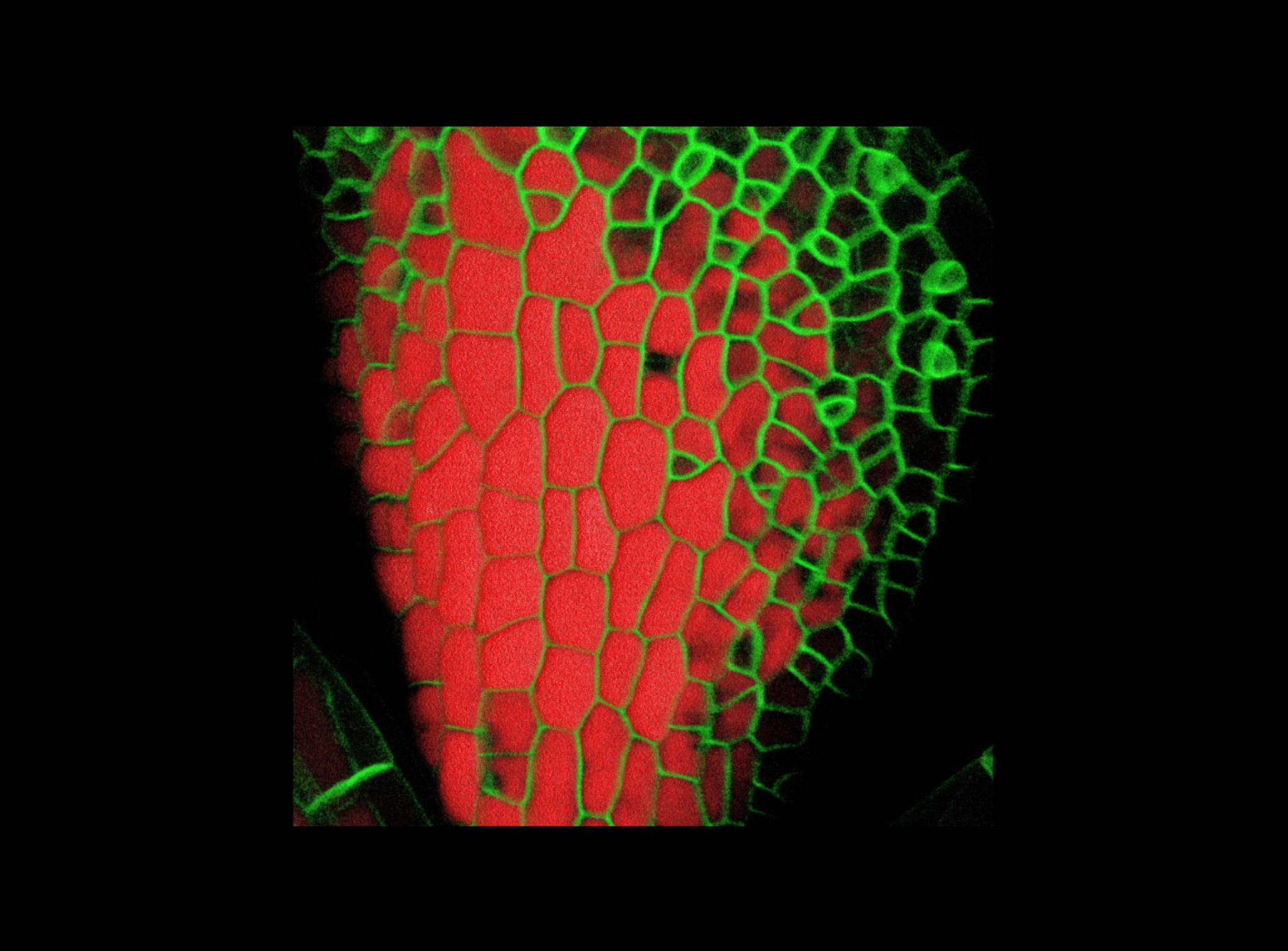
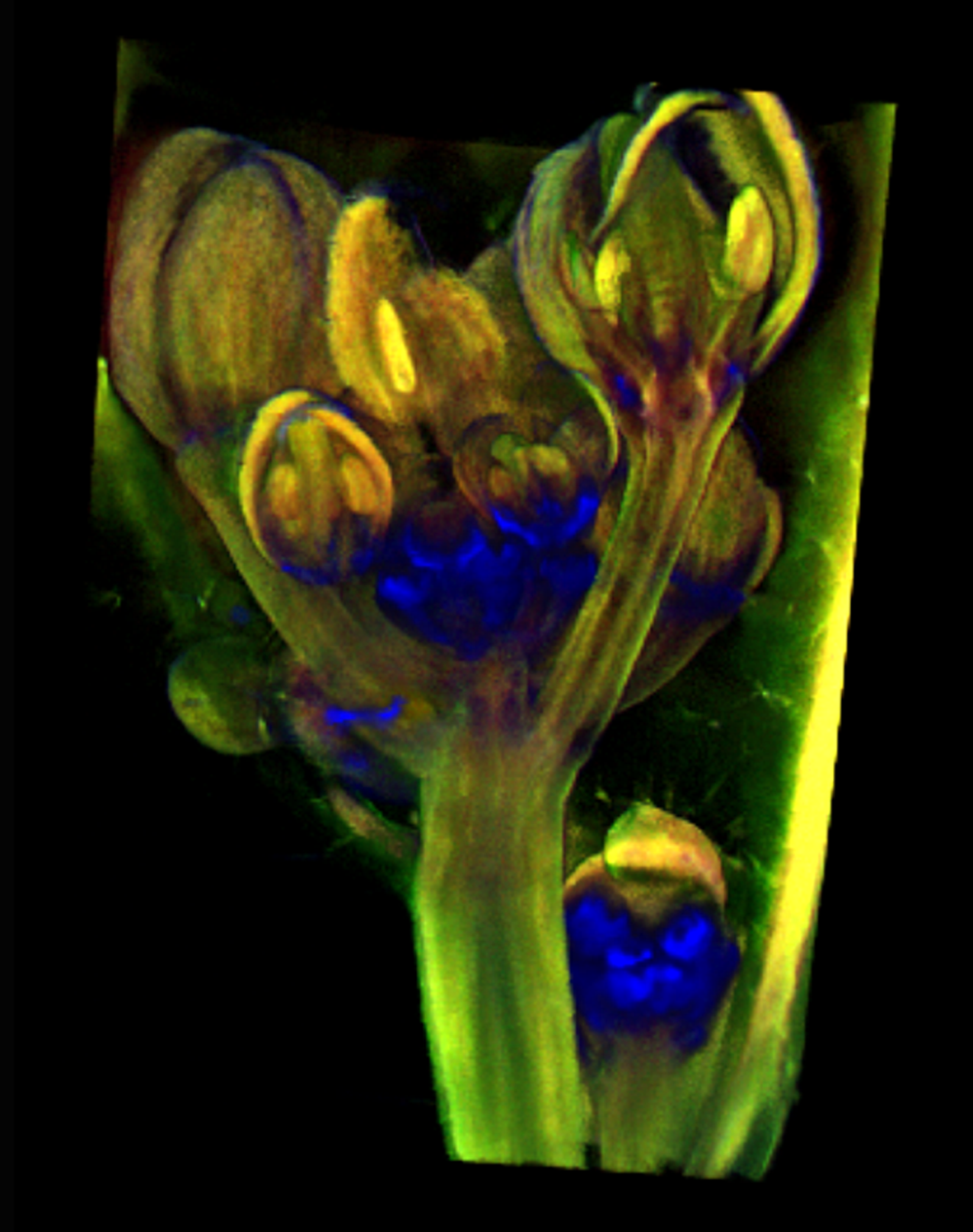
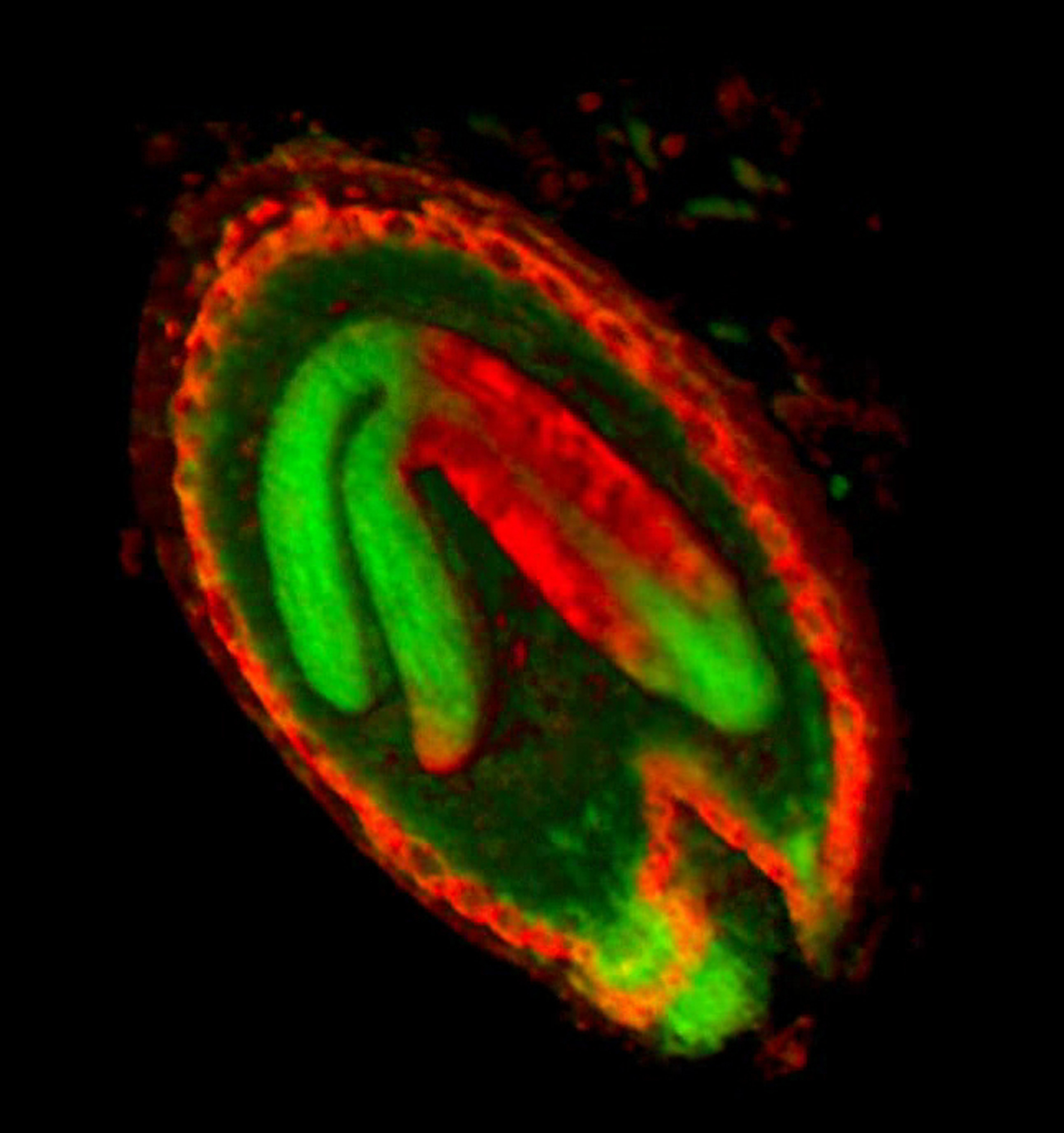
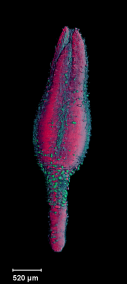
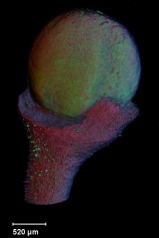
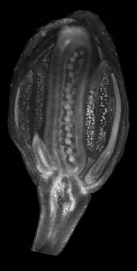
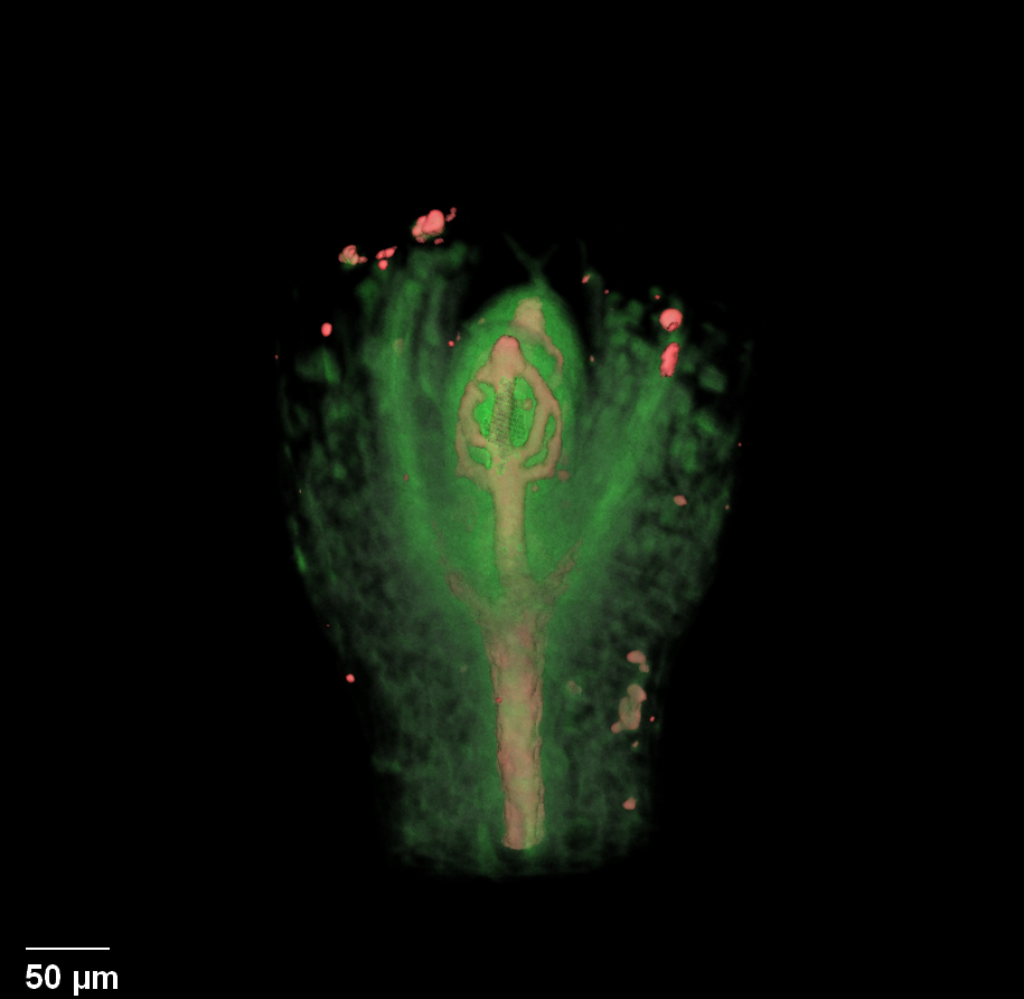
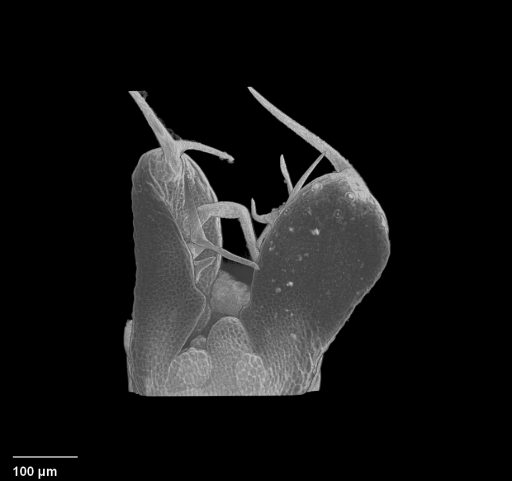
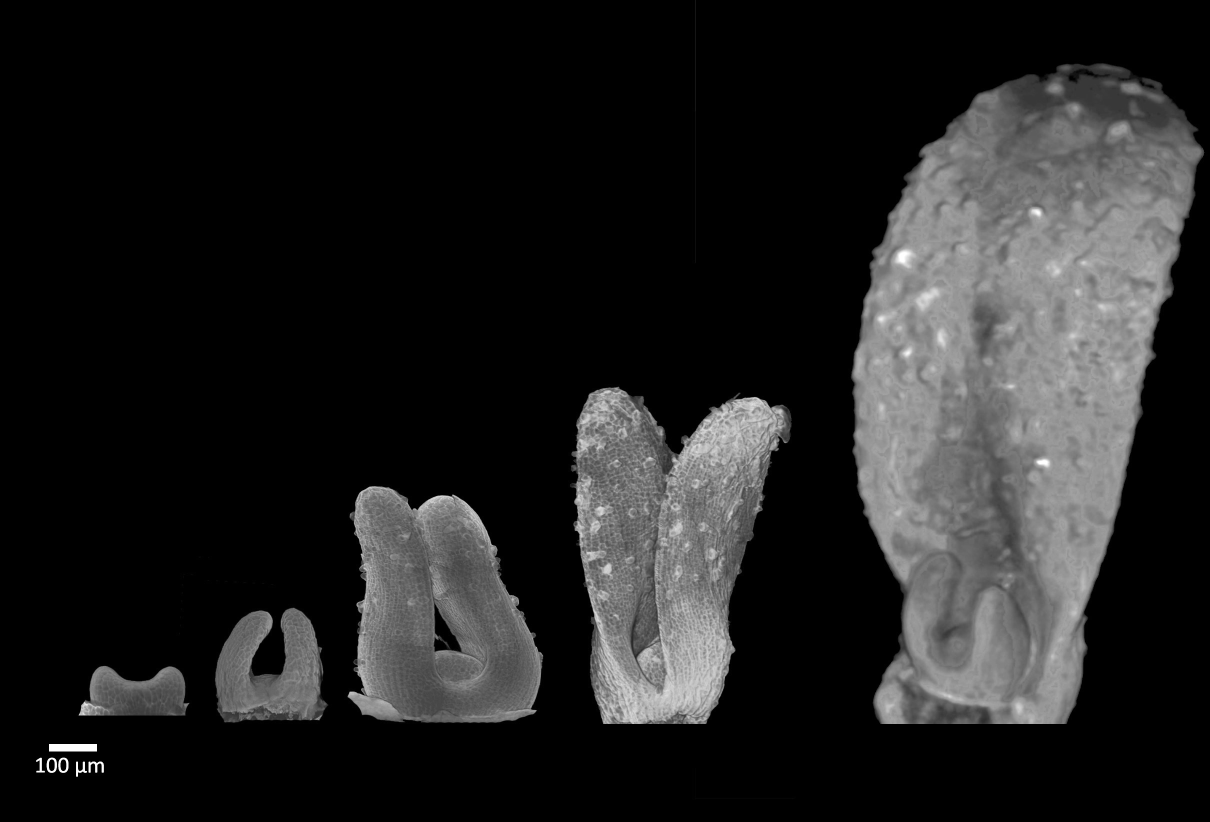
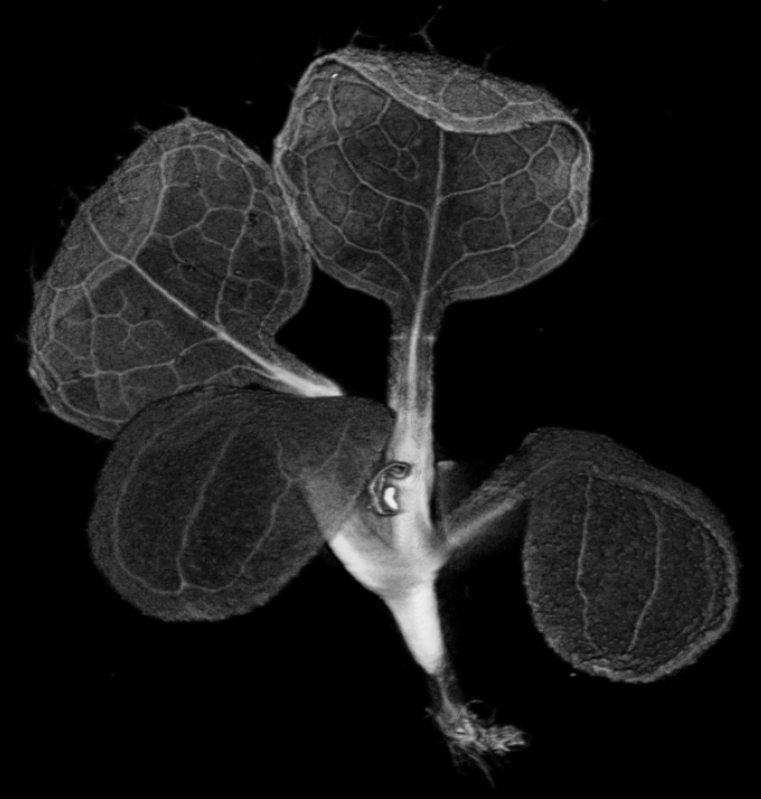
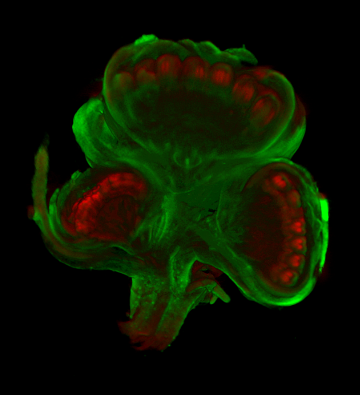
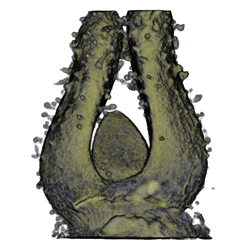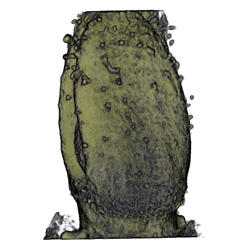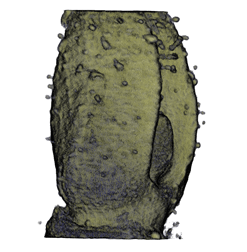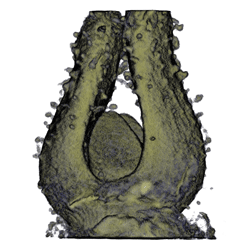
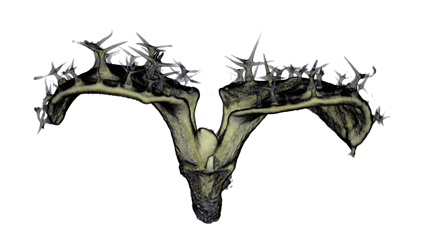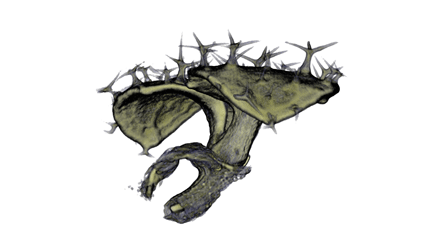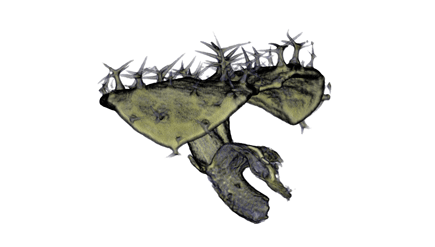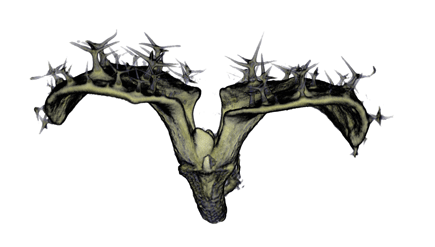
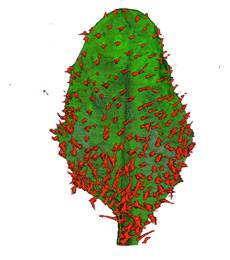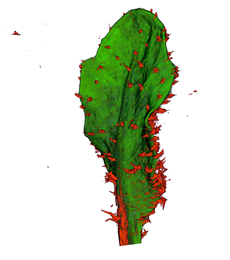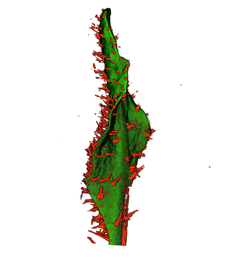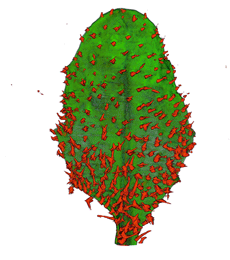
| Antirinhium Meristem | Arabidopsis Seedling | Arabidopsis Leaf (GL2:GUS expression in red) | Arabidopsis Leaf (Ath8:::GUS expression in red) |
| Download | Download | Download | Download |
* all data courtesy of Karen Lee [1]
Download
Although we try to keep up to date builds these sometimes lag behind the SVN trunk. So if you want the latest version / features, it is best to build the application from the trunk of the SVN. The build system is based on qmake for easy cross platform compilation.
OpenGL + Qt + OpenCL + LibTIFF + OMERO
| Windows (32bit) | Windows (64bit) | Linux | MacOS X (i386/x86_64/10.5+) |
OpenGL + Qt + LibTIFF
- Coming soon.
Windows Specific Notes
- You may need to install the corresponding Microsoft Visual C++ 2008 Redistributable Package which can be found here: 32bit and 64bit.
- WindowsXP users will need to change the view_gldrawbuffer = "GL_FRONT_AND_BACK" to view_gldrawbuffer = "GL_BACK" in the settings.ini file.
- The binaries are built with OMERO 4.3.4 support.
Source Code
Public SVN: https://cmpdartsvr3.cmp.uea.ac.uk/banghamlabSVN/VolViewer/
Building from source
Image Gallery
Media/Press
VolViewer has appeared in the following:
Cell: Online Gallery | Front cover: Handbook of Plant Science | Front cover: The Plant Cell | American Scientist | Royal Microscopical Society: Infocus Magazine | Bundled with the Bioptonic 3001 scanner: Bioptonics Viewer | The Daily Mail | The Guardian newspaper: 3D Fruit fly | Qt Ambassador program | Triffid Nurseries website
Author
- Dr Jerome Avondo Supported by the BBSRC through UEA Computing School and JIC.