VolViewer: Difference between revisions
JeromeAvondo (talk | contribs) No edit summary |
JeromeAvondo (talk | contribs) No edit summary |
||
| Line 21: | Line 21: | ||
<span style="color: Navy">'''How'''?</span> It is open source and written in C++ using OpenGL and Qt.<br> | <span style="color: Navy">'''How'''?</span> It is open source and written in C++ using OpenGL, OpenCL and Qt.<br> | ||
<span style="color: Navy">'''Where'''? </span>Binaries are available for the | <span style="color: Navy">'''Where'''? </span>Binaries are available for the Windows, Mac OS X and Linux, see below. | ||
Requirements: An OpenGL 2.1 / GLSL 1.20 compatible GPU with a recomended 512MB of memory. | Requirements: An OpenGL 2.1 / GLSL 1.20 compatible GPU with a recomended 512MB of memory. | ||
Revision as of 13:43, 5 March 2012
What? How? Where?
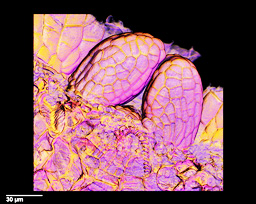
What? VolViewer is used for viewing volume images from, for example, confocal microscopy or optical projection tomography (OPT).
Features:
- Real-time volume rendering using an optimized 3D texture slicing algorithm.
- Interactive transfer functions to independently adjust opacity and intensity for up to three data channels.
- Real-time per channel thresholding, brightness and contrast operators.
- On-the-fly gradient computation for local illumination.
- Iso-surface computation with surface smoothing.
- Section viewing in any orientation / position.
- Real-time volume clipping.
- 3D measurements, filters & segmentation.
- Key frame interpolation for movie export.
- Stereo rendering using either quad buffer or anaglyph mode.
- Scripting interface to other systems, e.g. Matlab, OMERO, etc.
How? It is open source and written in C++ using OpenGL, OpenCL and Qt.
Where? Binaries are available for the Windows, Mac OS X and Linux, see below.
Requirements: An OpenGL 2.1 / GLSL 1.20 compatible GPU with a recomended 512MB of memory.
User Documentation
Sample Data
 |
 |
 |

|
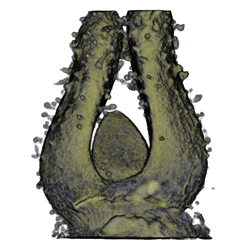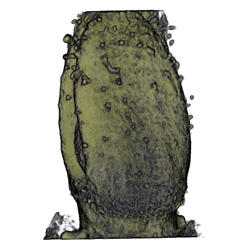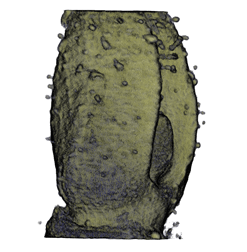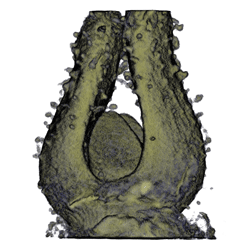
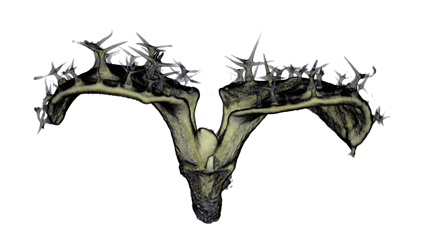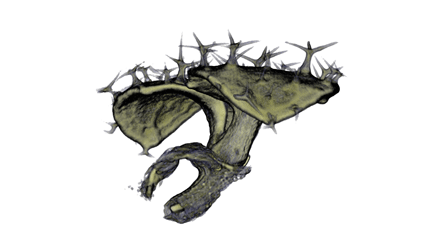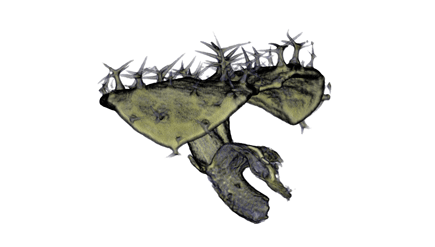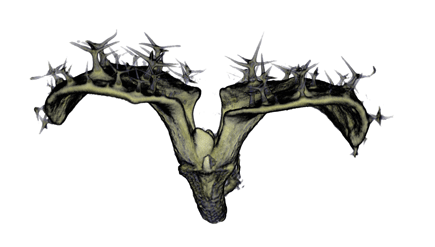
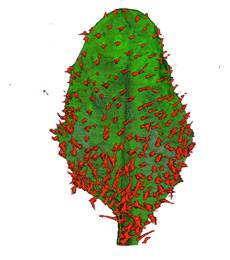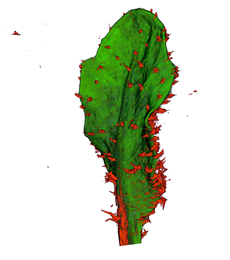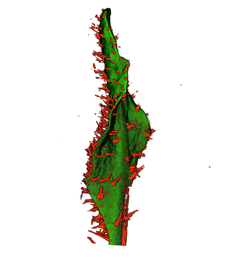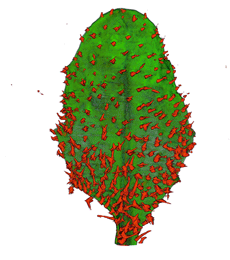
| Antirinhium Meristem | Arabidopsis Seedling | Arabidopsis Leaf (GL2:GUS expression in red) | Arabidopsis Leaf (Ath8:::GUS expression in red) |
| Download | Download | Download | Download |
* all data courtesy of Karen Lee [1]
Download
Although we try to keep up to date builds these sometimes lag behind the SVN trunk. So if you want the latest version / features, it is best to build the application from the trunk of the SVN. The build system is based on qmake for easy cross platform compilation.
| Windows (32bit) | Windows (64bit) | Linux | MacOS X (i386/x86_64/10.5+) |
Note that for the windows versions you will need to install the corresponding Microsoft Visual C++ 2010 SP1 Redistributable Package which can be found here: 32bit and 64bit.
Source Code
Public SVN: https://cmpdartsvr1.cmp.uea.ac.uk/banghamlabSVN/VolViewer/
Building from source
Media/Press
VolViewer has appeared in the following:
Front cover: Handbook of Plant Science | Front cover: The Plant Cell | Royal Microscopical Society: Infocus Magazine | Bundled with the Bioptonic 3001 scanner: Bioptonics Viewer | The Guardian newspaper: 3D Fruit fly | Qt Ambassador program | Triffid Nurseries website
Author
- Dr Jerome Avondo Supported by the BBSRC through UEA Computing School and JIC.